Mne bids pipeline: Difference between revisions
No edit summary |
|||
| (36 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
==Skip Background== |
|||
[https://megcore.nih.gov/index.php?title=Mne_bids_pipeline#Use_on_biowulf '''Use on Biowulf'''] |
|||
==Intro== |
==Intro== |
||
BIDS is a standard specification for neuroimaging/physiology data. This currently includes at least: MRI, fMRI, DTI, EEG, MEG, fNIRS (and possibly ECOG/sEEG). BIDS typically describes how RAW data is organized - and processed data is located in the bids_dir/derivatives/{AnalysisPackage}/{SUBJECT}/... The main advantage is that common code can be generated to process data organized in a standard format. Therefore, you should be able to import the bids data into any number of neurophysiological packages (MNE, Brainstorm, SPM, Fieldtrip, ...). Additionally, standardized processing packages known as BIDS apps can be used to process the data in the same way as long as the data is organized in BIDS. |
BIDS is a standard specification for neuroimaging/physiology data. This currently includes at least: MRI, fMRI, DTI, EEG, MEG, fNIRS (and possibly ECOG/sEEG). BIDS typically describes how RAW data is organized - and processed data is located in the bids_dir/derivatives/{AnalysisPackage}/{SUBJECT}/... The main advantage is that common code can be generated to process data organized in a standard format. Therefore, you should be able to import the bids data into any number of neurophysiological packages (MNE, Brainstorm, SPM, Fieldtrip, ...). Additionally, standardized processing packages known as BIDS apps can be used to process the data in the same way as long as the data is organized in BIDS. |
||
| Line 8: | Line 8: | ||
===BIDS website=== |
===BIDS website=== |
||
https://bids-specification.readthedocs.io/en/stable/ |
https://bids-specification.readthedocs.io/en/stable/ |
||
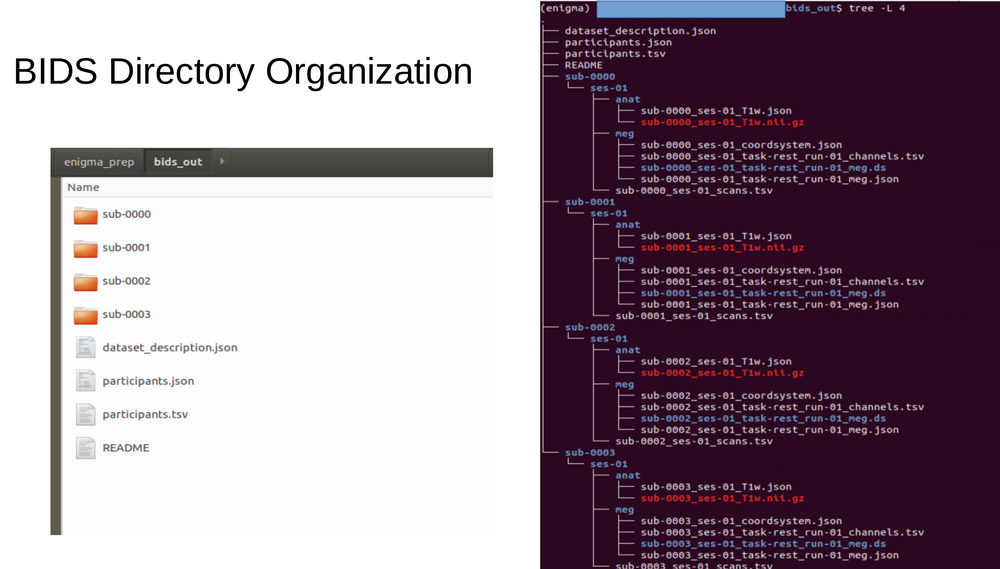
===Bids data organization=== |
|||
[[File:BIDS_data_organization.png|1000px|Bids data organization]] |
|||
==MNE Bids== |
==MNE Bids== |
||
| Line 27: | Line 30: | ||
https://github.com/mne-tools/mne-bids-pipeline/blob/main/config.py |
https://github.com/mne-tools/mne-bids-pipeline/blob/main/config.py |
||
===Example Config |
===Example Config=== |
||
(copy into a config.py text file and use) |
|||
Modify for use with your data - at a minimum: bids_root, conditions |
|||
study_name = 'TESTSTudy' |
study_name = 'TESTSTudy' |
||
bids_root = 'YOUR_BIDS_DIR' #<< Modify |
bids_root = 'YOUR_BIDS_DIR' #<< Modify |
||
| Line 35: | Line 41: | ||
epochs_tmax = 0.2 |
epochs_tmax = 0.2 |
||
baseline = (-0.1, 0.0) |
baseline = (-0.1, 0.0) |
||
resample_sfreq = 300.0 |
|||
ch_types = ['meg'] |
ch_types = ['meg'] |
||
conditions = ['stim'] |
conditions = ['stim'] #<< list of conditions |
||
| ⚫ | |||
#crop_runs = [0, 900] #Can be useful for long tasks that are ended early |
|||
| ⚫ | |||
==Use on biowulf== |
==Use on biowulf== |
||
!!If you run into any issues - please let Jeff Stout know so that the code can be updated!! |
|||
| ⚫ | |||
===Make MEG modules accessible=== |
|||
| ⚫ | |||
#Run the line below every login |
|||
#Or add the following line to your ${HOME}/.bashrc (you will need to re-login or type bash to initialize) |
|||
module use --append /data/MEGmodules/modulefiles |
|||
| ⚫ | |||
| ⚫ | |||
sinteractive --mem=6G --cpus-per-task=4 --gres=lscratch:50 #adjust mem and cpus accordingly - subjects will run in parrallel |
|||
| ⚫ | |||
module load mne_scripts |
module load mne_scripts |
||
| ⚫ | |||
#Afni coregistered mri data |
|||
| ⚫ | |||
## OR ## |
|||
#Brainsight coregistered mri data |
|||
make_meg_bids.py -meg_input_dir ${MEGFOLDER} -mri_bsight ${MRI_BSIGHT}.nii -mri_bsight_elec ${EXPORTED_BRAINSIGHT}.txt |
|||
===Freesurfer Processing=== |
|||
| ⚫ | |||
If you have already processed your freesurfer data - just copy or link the freesurfer data(see below). You can also set the freesurfer directory in mne-bids-pipeline config file. Alternatively, you can have the pipieline calculate freesurfer as part of the processing. This will take time - adjust sinteractive session (see below). |
|||
====Already Processed - Copy Data==== |
|||
mkdir -p ${bids_dir}/derivatives/freesurfer/subjects #If this folder doesn't already exists |
|||
#Copy your subject from the freesurfer directory to the bids DERIVATIVES freesurfer directory |
|||
cp -R ${SUBJECTS_DIR}/${SUBJID} ${bids_dir}/derivatives/freesurfer/subjects |
|||
====Process Using MNE-BIDS-Pipeline==== |
|||
#your sinteractive session must include --time=24:00:00 so that the freesurfer processing doesn't time out |
|||
#add the following step to your mne_bids_pipeline processing |
|||
--steps=freesurfer |
|||
| ⚫ | |||
First make a config file: [https://megcore.nih.gov/index.php?title=Mne_bids_pipeline#Example_Config example] |
|||
module purge |
module purge |
||
module load mne_bids_pipeline |
module load mne_bids_pipeline |
||
mne-bids-pipeline-run.py --config=CONFIG.py |
mne-bids-pipeline-run.py --config=CONFIG.py |
||
#Optional Flags |
|||
#--steps=preprocessing,sensor,source,report or all (default) - Can be a list of steps or a single step |
|||
#--subject=SUBJECTID(without the sub- prefix) |
|||
==TEST DATA for biowulf== |
|||
This section provides some test data to analyze using mne-bids-pipeline. Feel free to adjust parameters in the config.py after you run through the analysis the first time. All of the analysis parameters are defined in the config.py provided. Additional config.py parameters can be found above in [https://megcore.nih.gov/index.php?title=Mne_bids_pipeline#Processing Processing]. |
|||
====Start Interactive Session==== |
|||
| ⚫ | |||
====Copy and untar the data into your folder==== |
|||
cp -R /vf/users/MEGmodules/modules/bids_example_data_airpuff.tar ./ |
|||
tar -xvf bids_example_data_airpuff.tar |
|||
#Add the bids_root to your config file |
|||
echo bids_root=\'$(pwd)/bids_example_data_airpuff\' >> $(pwd)/bids_example_data_airpuff/config.py |
|||
====Load module and process the data==== |
|||
module load mne_bids_pipeline |
|||
mne-bids-pipeline-run.py --config=$(pwd)/bids_example_data_airpuff/config.py |
|||
#Copy this path for the next step |
|||
echo $(pwd)/bids_example_data_airpuff/derivatives/mne-bids-pipeline/sub-ON02747/ses-01/meg/sub-ON02747_ses-01_task-airpuff_report.html |
|||
#In another terminal download your results from biowulf |
|||
scp ${USERNAME}@helix.nih.gov:${PathFromAbove} ./ |
|||
''Open sub-ON02747_ses-01_task-airpuff_report.html in an internet browser to view the report'' |
|||
Revision as of 15:32, 26 May 2022
Skip Background
Intro
BIDS is a standard specification for neuroimaging/physiology data. This currently includes at least: MRI, fMRI, DTI, EEG, MEG, fNIRS (and possibly ECOG/sEEG). BIDS typically describes how RAW data is organized - and processed data is located in the bids_dir/derivatives/{AnalysisPackage}/{SUBJECT}/... The main advantage is that common code can be generated to process data organized in a standard format. Therefore, you should be able to import the bids data into any number of neurophysiological packages (MNE, Brainstorm, SPM, Fieldtrip, ...). Additionally, standardized processing packages known as BIDS apps can be used to process the data in the same way as long as the data is organized in BIDS.
BIDS (formal specs) >> MNE_BIDS (python neurophysiological BIDS) >> MNE_BIDS_PIPELINE (structured processing in MNE python according to BIDS definition)
BIDS website
https://bids-specification.readthedocs.io/en/stable/
Bids data organization
MNE Bids
MNE bids is a python tool to make and access neurophysiological data. This tool is a branch of the MNE tools for neurophysiological analysis.
MNE Bids website
https://mne.tools/mne-bids/stable/index.html
MNE bids pipeline background
Mne bids pipeline website:
https://mne.tools/mne-bids-pipeline/
Full example:
https://mne.tools/mne-bids-pipeline/examples/ds000248.html
Processing
All processing is defined in the config.py file. This has hundreds of options to define the processing.
Here are some of the typical options: https://mne.tools/mne-bids-pipeline/settings/general.html
Here is the full list of options:
https://github.com/mne-tools/mne-bids-pipeline/blob/main/config.py
Example Config
(copy into a config.py text file and use)
Modify for use with your data - at a minimum: bids_root, conditions
study_name = 'TESTSTudy' bids_root = 'YOUR_BIDS_DIR' #<< Modify l_freq = 1.0 h_freq = 100. epochs_tmin = -0.1 epochs_tmax = 0.2 baseline = (-0.1, 0.0) resample_sfreq = 300.0 ch_types = ['meg'] conditions = ['stim'] #<< list of conditions N_JOBS=4 #crop_runs = [0, 900] #Can be useful for long tasks that are ended early
Use on biowulf
!!If you run into any issues - please let Jeff Stout know so that the code can be updated!!
Make MEG modules accessible
#Run the line below every login
#Or add the following line to your ${HOME}/.bashrc (you will need to re-login or type bash to initialize)
module use --append /data/MEGmodules/modulefiles
Start interactive session with scratch to render visualization offscreen
sinteractive --mem=6G --cpus-per-task=4 --gres=lscratch:50 #adjust mem and cpus accordingly - subjects will run in parrallel
Create BIDS data from MEG data
module load mne_scripts
#Afni coregistered mri data
make_meg_bids.py -meg_input_dir ${MEGFOLDER} -mri_brik ${AFNI_COREGED}+orig.BRIK
## OR ##
#Brainsight coregistered mri data
make_meg_bids.py -meg_input_dir ${MEGFOLDER} -mri_bsight ${MRI_BSIGHT}.nii -mri_bsight_elec ${EXPORTED_BRAINSIGHT}.txt
Freesurfer Processing
If you have already processed your freesurfer data - just copy or link the freesurfer data(see below). You can also set the freesurfer directory in mne-bids-pipeline config file. Alternatively, you can have the pipieline calculate freesurfer as part of the processing. This will take time - adjust sinteractive session (see below).
Already Processed - Copy Data
mkdir -p ${bids_dir}/derivatives/freesurfer/subjects #If this folder doesn't already exists
#Copy your subject from the freesurfer directory to the bids DERIVATIVES freesurfer directory
cp -R ${SUBJECTS_DIR}/${SUBJID} ${bids_dir}/derivatives/freesurfer/subjects
Process Using MNE-BIDS-Pipeline
#your sinteractive session must include --time=24:00:00 so that the freesurfer processing doesn't time out #add the following step to your mne_bids_pipeline processing --steps=freesurfer
Process data using MNE Bids Pipeline
First make a config file: example
module purge
module load mne_bids_pipeline
mne-bids-pipeline-run.py --config=CONFIG.py
#Optional Flags
#--steps=preprocessing,sensor,source,report or all (default) - Can be a list of steps or a single step
#--subject=SUBJECTID(without the sub- prefix)
TEST DATA for biowulf
This section provides some test data to analyze using mne-bids-pipeline. Feel free to adjust parameters in the config.py after you run through the analysis the first time. All of the analysis parameters are defined in the config.py provided. Additional config.py parameters can be found above in Processing.
Start Interactive Session
sinteractive --mem=6G --cpus-per-task=4 --gres=lscratch:50
Copy and untar the data into your folder
cp -R /vf/users/MEGmodules/modules/bids_example_data_airpuff.tar ./ tar -xvf bids_example_data_airpuff.tar #Add the bids_root to your config file echo bids_root=\'$(pwd)/bids_example_data_airpuff\' >> $(pwd)/bids_example_data_airpuff/config.py
Load module and process the data
module load mne_bids_pipeline mne-bids-pipeline-run.py --config=$(pwd)/bids_example_data_airpuff/config.py
#Copy this path for the next step echo $(pwd)/bids_example_data_airpuff/derivatives/mne-bids-pipeline/sub-ON02747/ses-01/meg/sub-ON02747_ses-01_task-airpuff_report.html
#In another terminal download your results from biowulf
scp ${USERNAME}@helix.nih.gov:${PathFromAbove} ./
Open sub-ON02747_ses-01_task-airpuff_report.html in an internet browser to view the report