Brainsight Coregistration
BRAINSIGHT COREGISTRATION QUICK-START GUIDE [UNDER CONSTRUCTION]
OVERVIEW
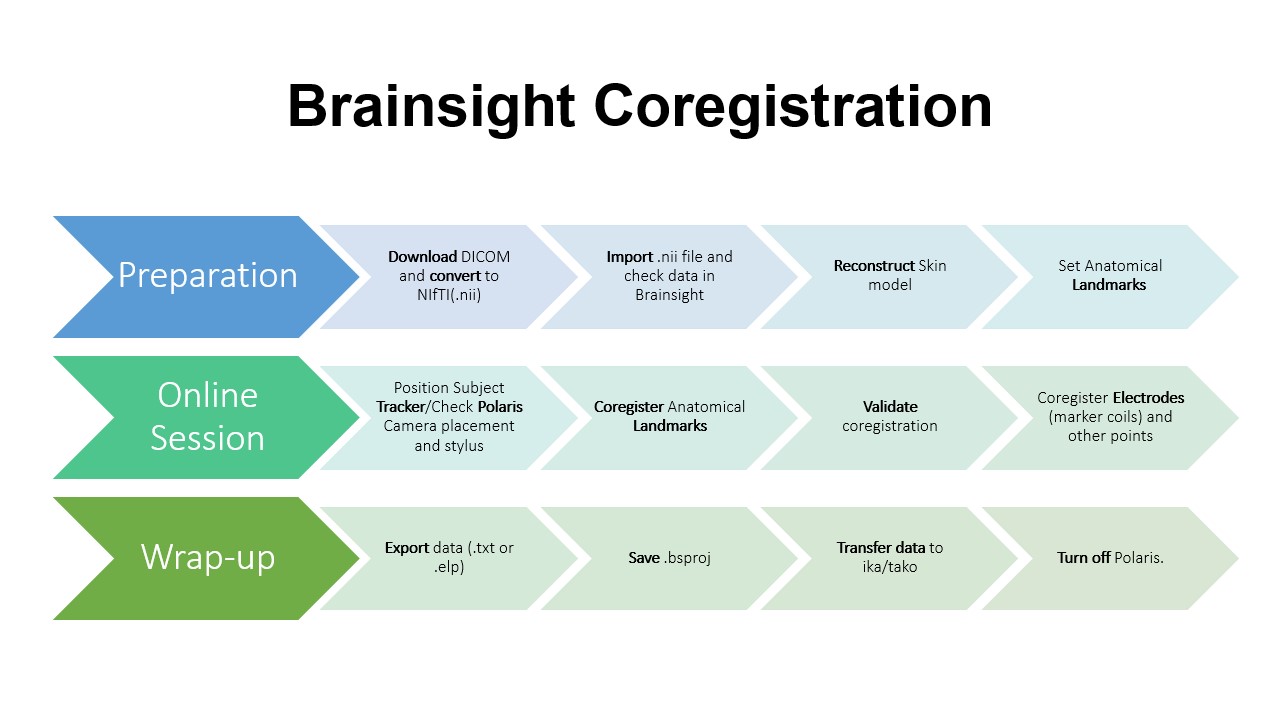
PREPARATION
Note: You will need your subject’s MRI data before your MEG appointment.
Downloading/Preparing the MRI data
If your participant’s data was collected here, you can download their data from the FMRIF DICOM Archive:
https://gold.nimh.nih.gov/
If you don’t have access to the archive – talk to your PI.
Transform the DICOM to NIfTI (.nii) format
- Locate the MP RAGE
The README file from the session will tell you which directory to use. If your subject doesn’t have an MPRAGE, you can use a suitable alternative (i.e. SPGR). If you’re unsure, consult your PI or MEG Core staff.
- Navigate to that directory in terminal and use the following command:
Dimon -infile_prefix anat_t1w_mp_rage_1mm_abcd -GERT_Rico -gert_write_as_nifti –gert_create_dataset
(If you use something other than MPRAGE, replace the prefix ‘anat_t1w_mp_rage_1mm_abcd’ accordingly.)
For more information on Dimon-- https://afni.nimh.nih.gov/pub/dist/doc/program_help/Dimon.html
- Generate MEG Hashcode for your subject
If you don’t have the opportunity to do it before your study, label your file something generic (I.e. SubjectXDate.nii). You will be able to rename it later; the primary goal is to remove the PII that NMRIF includes in the filename.
PII may still be buried in the header text of the data; treat the files accordingly.
Transferring data to the Brainsight Mac
- Transfer your data to your directory on tako
- Transfer your data from tako to one of Linux computers in the MEG Lab (namako or kani)
scp -r [SOURCE]@tako.nimh.nih.gov:/tmp/[MRI data] [DESTINATION]
- Using the dedicated MEG Lab flash drive, transfer your data to the Mac and file it in the appropriate MEGLAB directory for your study
BRAINSIGHT COREGISTRATION
Importing data (May be done before Subject arrives)
- Launch the Brainsight program and select “New Empty Project”. Note: If you do not have access to your subject’s MRI, choose “New MNI Head Project” and follow the Brainsight User Manual instructions for using an average brain template (see chapter 6). This is generally not recommended for our analyses
- Select subject’s anatomical data. NIfTI files are recommended by MEG Core, but other accepted formats are MINC, Analyze, DICOM, PAR/REC, or BrainVoyager VMR.
- When the data is loaded, select “Show Image and Details” to view the metadata and image volume, or proceed to the next step.
Automatic Skin Reconstruction
- From the 3D Reconstruction tab, select “Skin” from the “New” drop-down menu.
- If needed, adjust threshold and select “keep largest piece” to isolate the head from any MR noise
Setting anatomical landmarks
Note: These are anatomical landmarks clearly identifiable from the MRI. You are not placing the marker coils at this step.
- From the “Landmarks” tab, select “Configure Landmarks”. The landmarks window will display the Skin reconstruction and coronal, sagittal, and transverse slices. The cursor cross-hairs will be displayed in all four image windows.
- With the cursor cross-hairs identify the Nasion, LPA (left preauricular), and RPA (right preauricular). Rather than using the tragus, we will use the supratrageal notch -- notch above the tragus where the tragus joins the upper ear. This point is less likely to be distorted in the MRI by earplugs or head pads.
- For additional precision, you can select a fourth point on the tip of the nose or the outer canthus. You may use whatever point as long as it is unambiguously identifiable and fixed.
- Now you are ready to start a new session. At this point, you can suspend your project or move directly to the registration session.
Subject Preparation
Draw the marker coil locations on the subject.
1.5cm above the nasion, 1.5cm from the tragus on a line with the outer canthus of the eye.
Give the participant a skull cap and place the subject tracker headband on their head.
For EEG subjects who have already been gelled, use a green surgical cap.
Position them in the chair and select “New Online Session” from the session window. To select an existing session, highlight the desired session and select “Resume Session”.
Registration Session
With the Session window open, go through the following tabs(steps):
- Targets (optional). The anatomical landmarks selected can be imported here, but this section was intended for other applications (i.e. TMS, surgery).
- Polaris. Ensure that the Polaris Vicra is on and operational. We recommend turning on the Polaris as soon as you arrive so it has time to warm up.
Troubleshooting:
1. Polaris is too cold:
Leave the machine on for a few moments and refresh.
2. Subject tracker or stylus is not seen in the field of view:
Move the subject’s chair until they are seen.
Adjust the height of the camera stand.
3. All else:
Refresh unit or restart Brainsight.
If that fails: turn off unit, wait 30 seconds, restart.
Registration
Holding the stylus in both hands for stability, mark the anatomical landmarks selected in a previous step.
Click the “Speech Recognition” or “I/OBox Switch” option boxes to sample independently.
Identify the landmark with the stylus
Holding the stylus steady, say “SAMPLE” or step on the foot pedal.
If you have an assistant, they can click “Sample & Go to Next Landmark” for you, and give you feedback on the camera view (green or red) for the subject tracker and stylus.
Polaris will give you feedback
Successful: Polaris will chime and Brainsight will announce the next location to sample.
Unsuccessful: Polaris will give a negative beep.
Make sure both the subject tracker and stylus are in view of the camera.
Rotate the stylus
Ask the subject to look in a different direction.
Validation
Revisit the landmark samples. Brainsight will tell you the distance between the landmark and the sample, and the landmark and the skin. You ideally want less than 3mm (measurement will be green) but less than 5mm(measurement will be orange) is acceptable. Otherwise the distance will be red.
Run the stylus across the scalp. The crosshairs should follow the skull without intruding into the skull or introducing distance above.
If the measurements are unacceptable, it is either because the landmarks were incorrectly set (possibly due to distortions in the MRI) or because of poor sampling.
Resample or reset the landmarks until the validation is acceptable. When the registration is accurate, proceed to the “Electrodes” tab.
Electrodes
Click “Add From...” and select “Cap Layout” from the drop-down menu. In the pop-up window, select the file you wish to use.
For most studies, you can use “MEGCoreStandardCapLayout.txt”. This is recommended, at least as a starting point, if you use the MEG Core preprocessing pipeline.
For EEG studies, you may prefer a file that includes your EEG cap layout so you can digitize your electrodes as well.
Alternatively, you can create your own layout. Minimally, for analysis you will need:
- Nasion
- Left Ear
- Right Ear
Holding the stylus with both hands, sample the marker coil locations and optional cloud points. Brainsight will give you feedback as before.
When all the electrode points are satisfactorily sampled, click “Export to File” and save under the subject’s MEG Hashcode.
Saving as (.txt) is recommended, and we have scripts for converting .txt to an AFNI tagset (.tag). LOCATOR file format (.elp) is supported, but requires a few additional steps. Ask the MEG Core staff if this is something you want to do.
Save the Brainsight project under the subject’s MEG Hashcode.
Post-registration
Prep subject for entering the MSR as usual (bathroom, demetal, apply marker coils, etc.)
Save registration date to flashdrive and transfer to end destination.
You can leave the Mac on hen shut down the Vicra via the upper rocker switch, and the whole unit via the lower rocker switch.