Analysis with Nipype: Difference between revisions
Jump to navigation
Jump to search
Content added Content deleted
No edit summary |
No edit summary |
||
| Line 4: | Line 4: | ||
[[Image:Nipype meg.png | 800px]] |
[[Image:Nipype meg.png | 800px]] |
||
===Nipype Resources=== |
|||
https://nipype.readthedocs.io/en/latest/ <br> |
|||
https://miykael.github.io/nipype_tutorial/ <br> |
|||
===Nipype configuration for Biowulf=== |
===Nipype configuration for Biowulf=== |
||
Revision as of 11:29, 24 January 2020
UNDER CONSTRUCTION
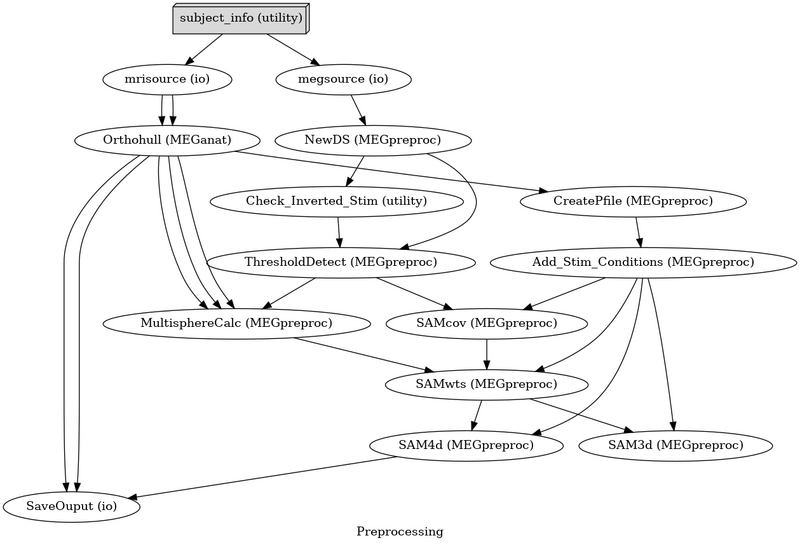
Nipype is a pipeline configuration tool that controls analysis flow and easily reconfigures for added functionality.
Nipye controls the flow of data and automatically determines dependencies to create filenames and folders
Nipype Resources
https://nipype.readthedocs.io/en/latest/
https://miykael.github.io/nipype_tutorial/
Nipype configuration for Biowulf
pull pyctf
module load fftw..... # first do a module spide fftw and find one to use Navigate to top folder (with Makefile) Make clean Make Make install Make usersite (will add a link so package is seen in conda)
pull SAMsrcV5
pull megblocks
install miniconda (or Anaconda)
- Add environment information to miniconda ##Need to provide environment.yml
- Add pyctf, SAMsrc, and megblocks to the miniconda environment ## TODO < provide a way to do this. Add the paths to miniconda/env......pth file
Nipype has strict requirements on inputs and outputs
- InputSpec
- OutputSpec
Preconfigured pipeline templates:
- Epilepsy
- Sam_epi
- Task
- Parsemarks, addmarks, samcov, sam3d, ... stats?
- Resting State
SAM input base class used for all of the SAM functions
Configuration on Biowulf:
- Set up conda: instructions on NIH HPC - https://hpc.nih.gov/apps/python.html
- Activate the conda environment and install nipype
#If you do not know your environment name type: conda env list conda activate $ENVIRONMENT_NAME conda install -c nipype conda-forge