Analysis AWS: Difference between revisions
No edit summary |
(→Done) |
||
| Line 53: | Line 53: | ||
'''Afni''' |
'''Afni''' |
||
afni -dset $MRI_file |
afni -dset $MRI_file |
||
==Done== |
|||
SAM v5 |
|||
singularity ? >> CTF |
|||
Afni |
|||
Jupyter |
|||
HNN |
|||
==TODO== |
==TODO== |
||
Revision as of 11:07, 7 November 2019
For Workshop members without access to computing resources - Preconfigured Amazon Web Services servers can be used
Accessing compute resources on AWS:
Speak with Jeff Stout to get the current IP address and PEM key
chmod 400 workshop.pem
${PATH} is the path to the PEM key provided to you
ssh -X -i ${PATH}/workshop.pem LOGIN_ID@IP_ADDRESS_PROVIDED
Windows Laptops do not have X11 installed natively. Download an Xterm software for visualization.
MobaXterm has been tested to work: https://mobaxterm.mobatek.net/download.html
Jupyter Notebook Over a Remote Connection
1) Run one terminal to start the notebook and forward the outputs to local web browser:
ssh -i AWS.pem -L 8887:localhost:8887 ubuntu@IP_ADDRESS conda activate workshop jupyter notebook --no-browser --port=8887
2) Copy the output of the jupyter notebook command (see example below) and enter into your laptop web browser (Firefox, Chrome, etc.). Do not click on the link - it will try to launch on the AWS computer
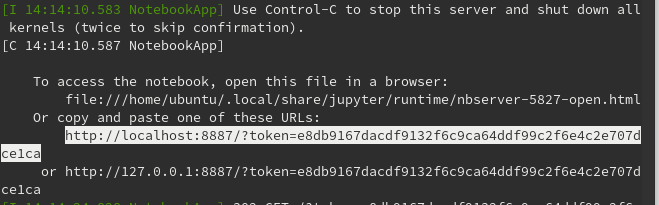
3) Use the jupyter notebook to analyze MEG data with python tools
Installed software
Human Neocortical Neurosolver
HNN website: https://hnn.brown.edu/
conda deactivate hnn
MNE python, Eelbrain, PyCTF
MNE website: https://mne.tools/stable/index.html
Eelbrain website: https://eelbrain.readthedocs.io/en/stable/
PyCTF website: https://megcore.nih.gov/index.php/MEG_Software_and_Analysis#MEG_Core_pyctf_tools_ported_to_Python_3
conda activate workshop # This conda environment has the MNE, Eelbrain, and PyCtf toolboxes installed ipython import eelbrain, mne, pyctf
CTF software
singularity shell /opt/ctf/ctf.img $ctf_command
singularity exec /opt/ctf/ctf.img $ctf_command
SAM version 5
sam_cov ... sam_3d ...
Afni
afni -dset $MRI_file
TODO
Freesurfer Reinstall MNE python Pyctf FSL Brainstorm Fieldtrip Spyder